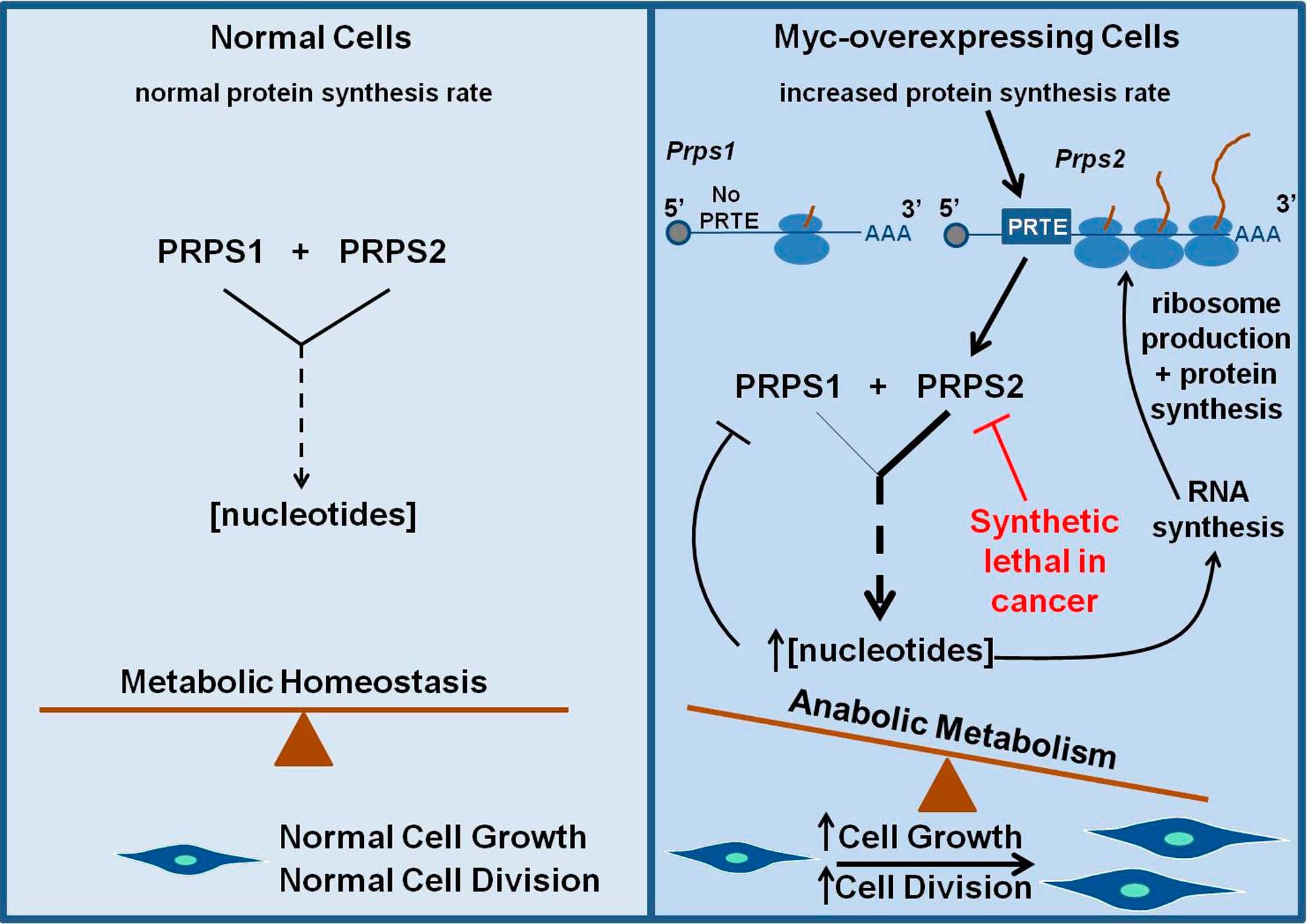
Cancer cells hijack important processes that provide constant energy (known as metabolism) and protein production (known as protein synthesis) to sustain their indefinite growth and proliferation. How cancer-causing genes, such as Myc, which is altered in almost all human cancers, control these increased energetic and protein production demands is an outstanding question that is still poorly understood.
In a study published in the May 22, 2014 issue of the prestigious scientific journal Cell, UCSF researchers led by Davide Ruggero, PhD, professor of urology, found that cancer cells exhibit a striking relationship between their ability to make more protein and the energy necessary to build the fundamental constituents that comprise a cancer cell. See here for article.
“We were able to use the unique tools we have developed in the lab to interrogate the relationship between protein synthesis and metabolism, which enabled us to identify a previously unrecognized connection between these two processes that, when altered, lead to cancer,” says Davide Ruggero, lead investigator of the study. “We hypothesized that the identification of this key connection could represent a point of vulnerability in cancer cells.”
The authors employed a candidate gene approach which revealed that a single enzyme, PRPS2 (phosphoribosyl-pyrophosphate synthetase 2), served as the mechanistic link tethering protein synthesis and metabolism. Surprisingly, the authors found that the Myc oncogene directly increases the amount of the PRPS2 enzyme, which was sufficient to regulate these two processes, leading to cancer cell growth.
“By regulating PRPS2 directly, cancer cells can quickly produce the cellular building blocks to promote their uncontrolled growth and division,” says Tom Cunningham, a postdoctoral fellow in the Ruggero lab and first author of the study.
Ruggero and his team then generated mice genetically manipulated to lack PRPS2 to test the enzyme’s function in normal physiology and cancer development. Strikingly, the PRPS2-deficient mice displayed no phenotypic differences compared to normal mice, however they were profoundly resistant to developing cancer.
“We believe the clinical implications of this study could be huge,” says Ruggero. “Because normal cells can tolerate loss of PRPS2 function, but cancer cells cannot, the therapeutic window for new drugs that target PRPS2 could be very wide. The identification of this metabolic enzyme as an Achilles’ Heel for cancer cells may also provide a means of therapeutically targeting cancers caused by the activation of currently “undruggable” oncogenes such as Myc.”
The researchers are now beginning to explore the possibility that the mechanism they have uncovered has broad application in many different human cancers.
“We’d like to see how far down the rabbit hole this discovery takes us,” says Ruggero. “If we see that a multitude of human cancers rely on the function of this metabolic enzyme for their continued proliferation and survival, then we may have a fantastic new avenue to impact patient outcomes in the clinic.”
These studies were performed in collaboration with postdoctoral researcher Alessia Lodi and Sabrina Ronen, PhD, professor of radiology. Melissa Moreno, a technician in the Ruggero Lab, also contributed to this work. Funding for this research came from the Leukemia and Lymphoma Society Scholar, America Cancer Society grant 121364-PF-11-184-01-TBG; and National Cancer Institute grants R01 CA154916; R01CA140456; R01 CA130812; and R01 CA154915.